Having extracted and assessed the total RNA from your samples, you now have the task of creating the first strand complementary DNA (cDNA). Sounds easy enough, right? Well, when it comes to creating cDNA there are multiple options available in terms of what type of primers to use in your reaction. These include random primers or oligo(dT)s, as well as other alternatives such as sequence-specific (also known as gene-specific) primers.
Below is a description of each option to help you decide which one is best for your purposes.
Mastering qPCR
A video tutorial explaining the different cDNA synthesis options can be found in our Mastering qPCR course
>>Use code 20QPCR to get 20% off<<
Random primers
Random primers are short oligonucleotides that have a random sequence. Since they are usually 6 nucleotides in length, they are often known as random hexamers. The random sequence enables the unbiased binding of primers to anywhere on most types of RNA including rRNA and small RNAs. Since they bind throughout, the majority of the gene products should be covered, therefore increasing the chance of amplification during PCR.
Oligo(dT) primers
Oligo(dT) primers are composed of stretches of deoxythymidine. They are available in various lengths, with the most common being 12-18 nucleotides long. Oligo(dT) primers are designed so that they bind to the complementary poly(A) tails of messenger RNA (mRNA). Therefore, oligo(dT)s are only useful in cDNA reactions when mRNAs are the target for the downstream application. They will not anneal to non-polyA mRNA fragments, such as 18S rRNA. Also, bear in mind that if your PCR primers are designed near the 5’ end of a gene, the use of oligo(dT) primers for cDNA synthesis is not the best idea. Since these primers anneal to the 3’ end of mRNA, the 5’ end may not be present in the extended product, especially if you are investigating a large gene.
Sequence-specific primers
Sequence-specific primers provide the greatest specificity for the target of interest since unique primers are used that will only convert the region of interest. The downside of this, however, is multiple cDNA reactions with different sequence-specific primers will be required if more than one region is to be targeted in the downstream PCR reaction.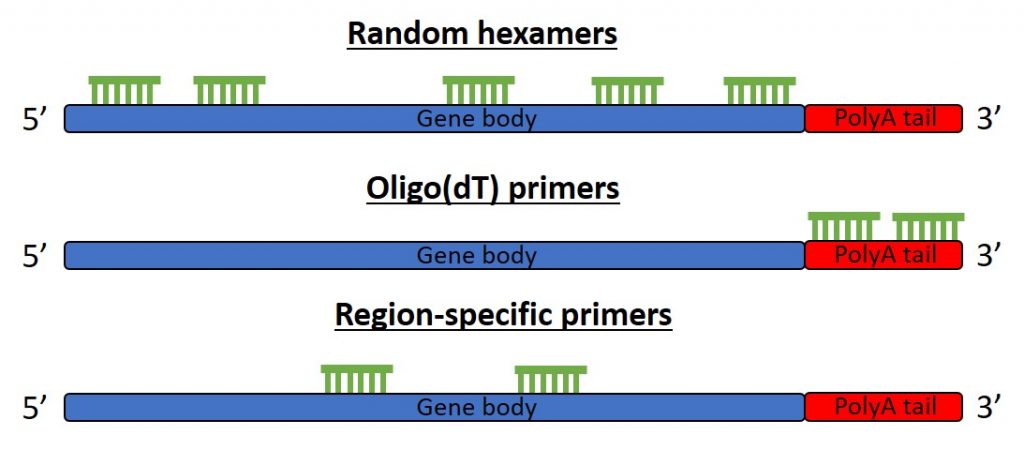
Which is the best option for cDNA synthesis prior to PCR?
The choice is usually between random hexamers and oligo(dT) primers. The decision will be based on your target of interest. It is no good using oligo(dT)s if you plan to amplify 18S rRNA, which is frequently used as a housekeeping gene.
The best advice, if you are looking at mRNA in PCR, is to use a mixture (50:50 split) of both random hexamers and oligo(dT) primers during cDNA synthesis.




Hi
This is very clear and helpful. Thanks so much
Thank you this was really helpful
Steven
Thank-you for the this concise and clear explanation.
Hey Steven,
Thanks so much for the clear explanation. well appreciated.
hi steven ,
does the random hexamer primer in RT step can be used for micro RNA detection ?