What is the Polymerase Chain Reaction (PCR)?
PCR is a molecular technique frequently applied to amplify regions of interest in DNA, resulting in millions or billions of copies.
The PCR reaction is made possible through the use of the thermostable DNA polymerase enzyme known as Taq polymerase. Taq refers to the heat-resistant bacteria (Thermus aquaticus) the polymerase is found. Using this heat-stable DNA polymerase enzyme is essential since the PCR reaction involves cyclic changes in temperature ranging from 50 – 98oC, therefore unlike other polymerase enzymes, this one will not degrade when exposed to extreme temperatures.
Components of the PCR
Performing PCR is relatively simple and cheap to do. All that is required are four components.
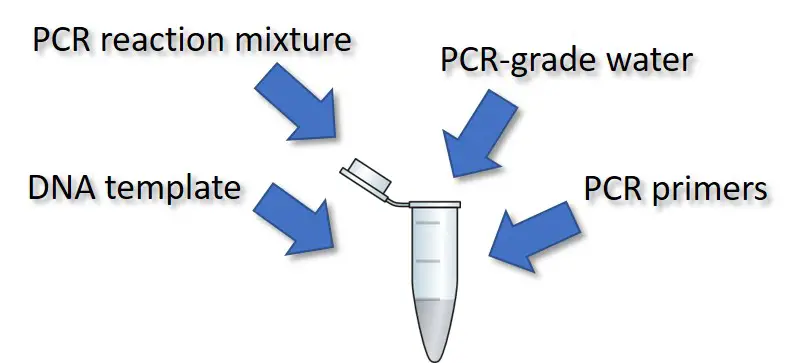 DNA template
DNA template
The DNA template is the starting material for the PCR. It contains the region you want to amplify. Examples of template DNA used in PCR include genomic DNA (gDNA) and complementary DNA (cDNA).
PCR primers
PCR primers are short pieces, usually around 20 bps in length, of single-stranded DNA. The primers are made as a pair: a forward and reverse primer, which are designed to bind to the complementary sequence surrounding the region you want to amplify.
PCR-grade water
PCR-grade water is pure water which is free from DNase and RNase enzymes. The water is used to normalise the total volume between the samples.
PCR reaction mixture
The PCR mixture contains the Taq DNA polymerase, the enzyme responsible for performing the amplification, nucleotides, as well as other substances required for the PCR reaction, such as magnesium ions. Sometimes these components are supplied separately.
Overview of the PCR procedure
A typical PCR procedure can be broken down into different steps. These are described in more detail below.
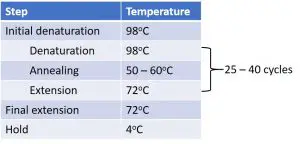
 1. Initial denaturation
1. Initial denaturation
This step is performed for two reasons. The first is to activate the Taq polymerase enzyme. The second is to denature the DNA starting material. The time used for the initial denaturation step will vary depending upon the type of Taq polymerase used and type of starting material. For example, genomic DNA requires longer incubation times since it is double-stranded and contains more bonds that have to be broken, whereas cDNA is already single stranded so requires less time.
2. Cycle phase
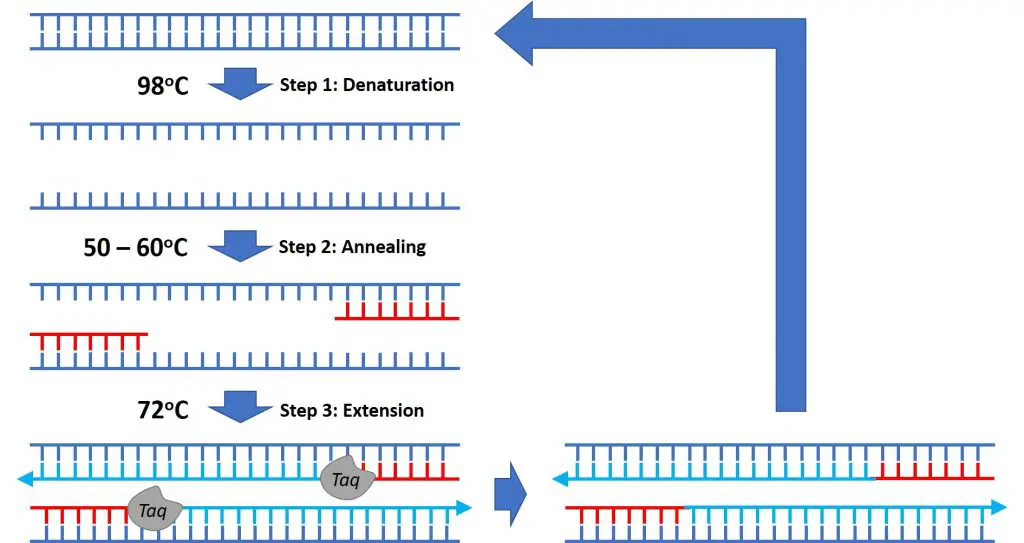 Now begins the start of the cycle phase containing a denaturation, annealing and extension step. This is the vital part of the PCR procedure in order to amplify the region of interest. Generally, the cycle phase is repeated 25 – 40 times for optimal amplification. After each cycle, the amount of DNA amplified will double, therefore, the larger the number of cycles used, the more material that will be amplified at the end of the reaction.
Now begins the start of the cycle phase containing a denaturation, annealing and extension step. This is the vital part of the PCR procedure in order to amplify the region of interest. Generally, the cycle phase is repeated 25 – 40 times for optimal amplification. After each cycle, the amount of DNA amplified will double, therefore, the larger the number of cycles used, the more material that will be amplified at the end of the reaction.
Cycle step 1: Denaturation
Like the initial denaturation, the purpose of the denaturation step in the cycle phase is to denature the template material. The material will be heat denatured from double-stranded DNA to single-stranded DNA to enable primer annealing.
Cycle step 2: Annealing
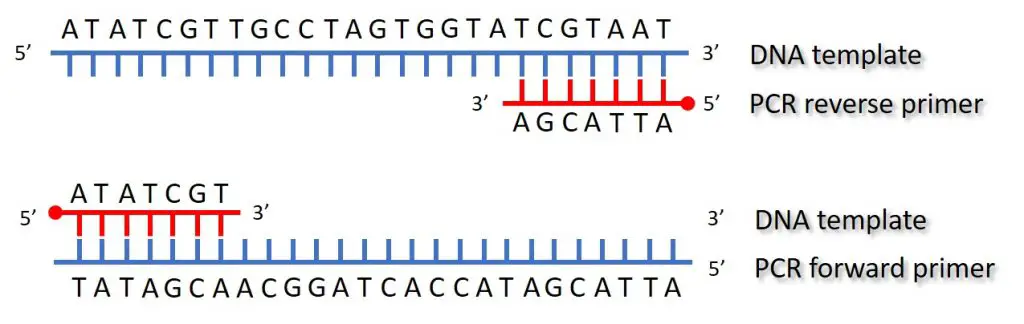 The temperature of the annealing step will vary depending upon the properties of the primers (such as the length and composition) used in the reaction. However, the lowering of temperature at this step is to enable primer binding to the single-stranded template.
The temperature of the annealing step will vary depending upon the properties of the primers (such as the length and composition) used in the reaction. However, the lowering of temperature at this step is to enable primer binding to the single-stranded template.
Primers binds to the complementary sequence, see the above image for a schematic of primers binding to the DNA template.
Cycle step 3: Extension
The temperature is then raised again to 72oC to enable the Taq polymerase to bind to the 3′ prime end of the primers and extend the sequence. Eventually, the enzyme will copy the complementary sequence of DNA, thus reforming a double-stranded template again.
The three cycle steps are then repeated, usually 25 to 40 times, to amplify the region of interest.
3. Final extension
Some protocols suggest adding a final extension step at the end of the cycle phase to ensure that all of the Taq polymerases have finished extending.
4. Hold
This is also an optional addition at the end of the procedure but is usually recommended, to basically keep the samples at a low enough temperature for storage and to avoid denaturation. This is particularly handy when you want to run a PCR program overnight and want to collect the samples the next day.