What is Protocols.io?
Have you ever been stuck in the lab for weeks on end attempting to perform a particular method and cannot understand why it isn’t working? Most of the time, protocols are passed down within labs like family heirlooms. I can personally say that this isn’t very efficient. Well, let me take a moment to introduce you to Protocols.io.
Protocols.io is an open-access protocol reprository for the life sciences. It has a virtual community platform to allow researchers to develop, discuss and share their perfectly optimised protocols. This makes reproducing techniques incredibly more time and cost effective.
As well as this, researchers are encourgaed to upload their detailed protocols to Protocols.io and refer to these when publishing their work within the journal article. An example of this is that of Yao and colleagues, as published in PLOS Biology. At the start of the materials and methods section, the authors supply the link to the corresponding Protocols.io folder:
 When you follow the link, a folder will open containing all of the relevant techniques and reagents used in the experiment via the Protocols.io platform. It is essentially like looking into their lab notebook to see exactly how they performed each experiment or method. This then allows others to replicate their findings. So simple, yet highly efficient!
When you follow the link, a folder will open containing all of the relevant techniques and reagents used in the experiment via the Protocols.io platform. It is essentially like looking into their lab notebook to see exactly how they performed each experiment or method. This then allows others to replicate their findings. So simple, yet highly efficient!
Listed below are some of the main features of the Protocols.io platform.
The features of Protocols.io
Upload and share your protocols
The primary feature of Protocols.io is the ability to upload and share your own protocols. When you first begin, a quick tour guide of the protocol editor to explain all of the functionality available to you. The editor itself is so easy to use that you can just dive straight in. You get the ability to add steps, each with their own title and body of text. You can also add components, such as a timers, safety information and kit information. Another nice inclusion is the ability to add a manuscript citation to a protocol if it is linked to a paper.
 If you like the look of a certain protocol, you can also make your own version of it by creating a ‘fork’. Here, you use the original protocol as a template while you adjust it to your own version.
If you like the look of a certain protocol, you can also make your own version of it by creating a ‘fork’. Here, you use the original protocol as a template while you adjust it to your own version.
All protocols are set to ‘Private’ by default, meaning they are not shared publicly on the Protocols.io repository. But you do have the option to publish them if you so wish, which I would encourage everyone to do. Other sharing options available include the ability to export protocols as a PDF and sending email links.
Run a protocol
A nice feature is the ability to ‘Run’ a protocol (this requires you to have an account with Protocols.io – which is free!). By running a protocol you can tick off the steps you have completed so far. If a step requires an incubation, it will also notify you and count you down. No need to find a timer! If you don’t have time to complete the whole run, you can also save your progress in your ‘Journal’ and continue it later on.
 Connect with other researchers
Connect with other researchers
When you create an account with Protocols.io, you get the ability to discuss with other users. Part of the site has a dedicated discussion forum with a questions and answers board.
Within each protocol, users also get the ability to post comments to the creator for further information. This feature works exceptionally well if the protocol is a little unclear in parts since the authors are already on Protocols.io, you usually get an answer without much delay.
If the creator of a protocol has added an annotations feature to certain steps, users can then post small comments to provide their expert opinion. For example, maybe instead of using the quoted 100 mL of a solution, you found that 80 mL gave better results. You can let others know by adding an annotation.
The mobile app
A big feature of Protocols.io is their mobile app. The app is available on both the iTunes and Google Play Store to support the majority of devices. Here you can quickly browse through protocols on the go.
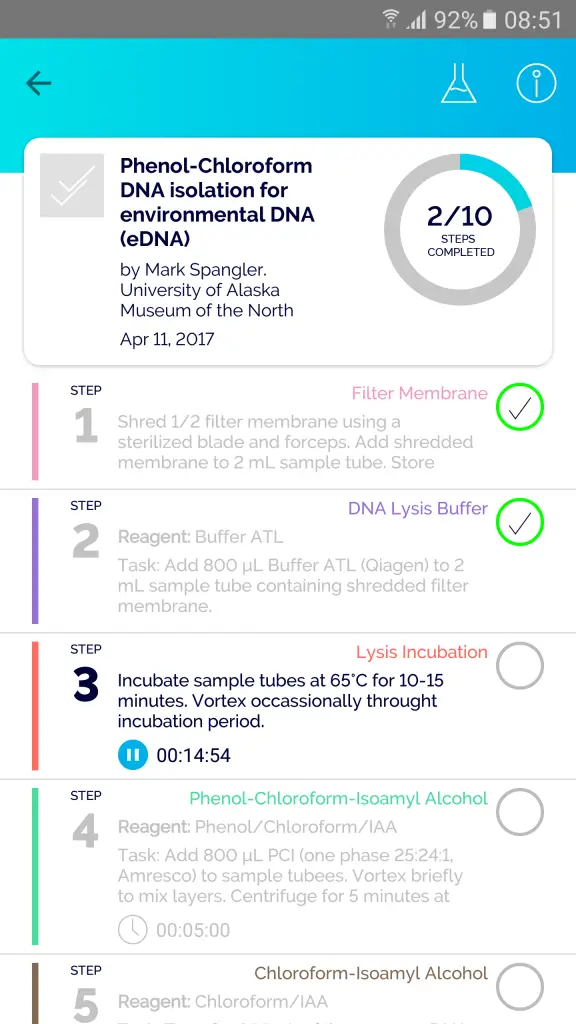 Other useful perks of the mobile app include; multiple timers (4 at the time of writing) to keep track of different steps, a molarity calculator for those tricky calculations if you don’t have time for the old C1V1 = C2V2 workings, and a journal where you can create project folders and have specific protocols saved in them.
Other useful perks of the mobile app include; multiple timers (4 at the time of writing) to keep track of different steps, a molarity calculator for those tricky calculations if you don’t have time for the old C1V1 = C2V2 workings, and a journal where you can create project folders and have specific protocols saved in them.
Widgets
If you have your own website or blog, you can insert a protocol as a widget. Widgets are usually small parts of hmtl code which you can incorporate into a website to add further functionality.
Here is an example of a widget created by New England Biolabs on Protocols.io:
Protocols.io is the future of protocol sharing. I encourage every researcher to adopt the use of the platform to complement their published work. This will ultimately help boost the confidence of research which has otherwise been struck by reproducibility concerns over recent times.