Gone are the days where you have to design primers by hand. Instead, we have a plethora of excellent resources available that utilises complex algorithms to determine the optimal primers for your PCR reaction. These give the best theoretical chance of your PCR reaction working, thus saving time and money. Best of all, the majority of them are free to use or download. Here is a list of 13 free primer design programs to use when designing primers:
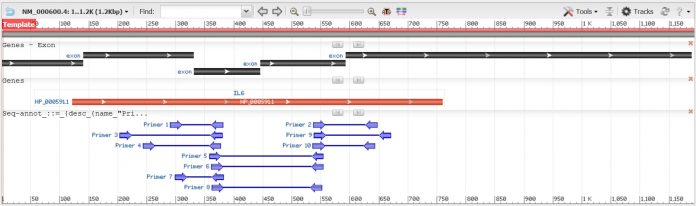
Primer-BLAST was developed by NCBI and utilises the Primer3 platform in combination with the BLAST algorithm. The incorporation of BLAST enables the screening of potential primers to unintended sequences in the organism’s genome, which is an extremely important feature. It also gives you an informative visual output for where the primers will bind on the target sequence. Primer-BLAST is by far my favourite and number one choice when designing primers. If you are thinking of using Primer-BLAST to design primers for PCR, be sure to check out my quide on how to do this.
Primer3 has a very minimalist feel, however, it has very powerful and comprehensive primer design capabilities. There are a wealth of parameters to tweak to find the best set of primers for your target. Because of this, some may find it quite overwhelming, so I would suggest the more advanced users take advantage of Primer3. If you prefer to use the standalone program, instead of the web interface, you can download it for free here.
Primer3Plus is another web-based primer design interface that is based on the Primer3 platform. It is a bit more user-friendly than Primer3 itself, with each primer parameter separated onto multiple tabs.
PrimerQuest is Integrated DNA Technologies (IDT) answer to successful primer design. You can chose to design primers based on a manually entered sequence, using an NCBI ID number (NM number, for example) or by uploading a sequences in an Excel file. The specific type of PCR primers (general PCR, qPCR with and without probes and sequencing) can also be selected for application-specific outputs.
OligoPerfect is Thermo Fisher Scientific’s primer design interface. Start by entering your ‘Sequence Name’, ‘Application (PCR:Detection)’ and ‘Researcher Name’. Then either enter your target sequence or import the sequence as a FASTA file. You then get the option to tweak the desired primer parameters. These parameters are not as in-depth as other primer design interfaces but may suffice for certain primers.
PerlPrimer is an open-source GUI program that is free to download. You have the ability to design primers for standard and qPCR. The application is very simplistic and is very easy to use. You also have the ability to BLAST the primers using the NCBI server, which is a must for any good primer design platform.
OLIGO is another primer design software that is free to download. It even comes with a PDF tutorial guide to help you through installing and using OLIGO. OLIGO also allows the ability to design TaqMan probes for qPCR.
The GenScript Real-time PCR (TaqMan) Primer Design online tool is specifically suited for TaqMan primer and probe creation. It is nice and simple to use with minimal primer parameters to set. You can customise the PCR amplicon size, primer and probe Tm and choose to have the primer/probe crossing an exon junction.
GenScript also have the GenScript Online PCR Primers Designs Tool for standard PCR primer design. You have more flexibility, compared to their Real-time PCR (TaqMan) Primer Design interface, in setting a plethora of primer parameters by using their advanced tab.
AutoPrime is specific for qPCR applications that want to measure eukaryotic gene expression. The platform is based on Primer3, and puts a lot of emphasis on amplifying cDNA, therefore only primers spanning exon-exon boundaries and those separated by an intron will be generated. You don’t even have to input a FASTA sequence, simply enter your gene name and you are good to go.
RExPrimer is a unique primer design application in that it will notify you of any structural variations in the target region, such as the presence of SNPs, insertion/deletion polymorphisms and pseudogenes. This will allow for more efficient primer binding and ultimately improve your PCR results.
BatchPrimer3 is yet another Primer3-based primer design software freely available online. There is a huge amount of primer subtypes to design including, generic PCR primers. BatchPrimer3 requires a FASTA sequence to be entered or uploaded. An intermediate selection of primer parameters are also there to tweak.
The Eurofins Genomics’ Primer Design Tools uses the Prime+ program from the GCG Wisconsin Package, originally written by Irv Edelman. All that is required is a target sequence, which is copied and pasted into the program, then you have the option to tweak the design parameters. The output also includes a suggested annealing temperature to use for each primer pair.
Featured image credit: MIKI Yoshihito (via Flickr)
Do you have any free PCR primer design software suggestions to add to the list? Leave a comment below with your preferred choice and I will be sure to update the list.




Thank you so much Dr Steven. Your article helped me a lot 🙂
Hi
Thank You Steven for the great help.
I need to know if any of these softwares can help me design multiplex pcr primer sets
I need something to help me analyse Self Complementary structures
Regards
Hi Nima
Many thanks for your comment.
For multiplex primer design tools that are free, your choices are rather limited. I did see a recent one called oli2go. I have never used this, but it does look like it may help you out.
If you find any more, please let me know and I can update the list.
Best wishes,
Steven
hi steven ,
thanks alot for ur incredible article , i wish you tell me which tool would be useful for divergent primer design ?