What is bisulfite pyrosequencing?
Bisulfite, also referred to as bisulphite, pyrosequencing is a technique used to quantify DNA methylation levels on DNA at a single-base resolution. In this article, I will explain how bisulfite pyrosequencing works.
DNA methylation is most commonly found to occur when a cytosine (C) is adjacent to a guanine (G) base, which is known as a CpG site. The ‘p’ denotes the phosphodiester bond which connects the two bases. Specifically, DNA methylation occurs on the C base of the CpG site at the 5th carbon position.
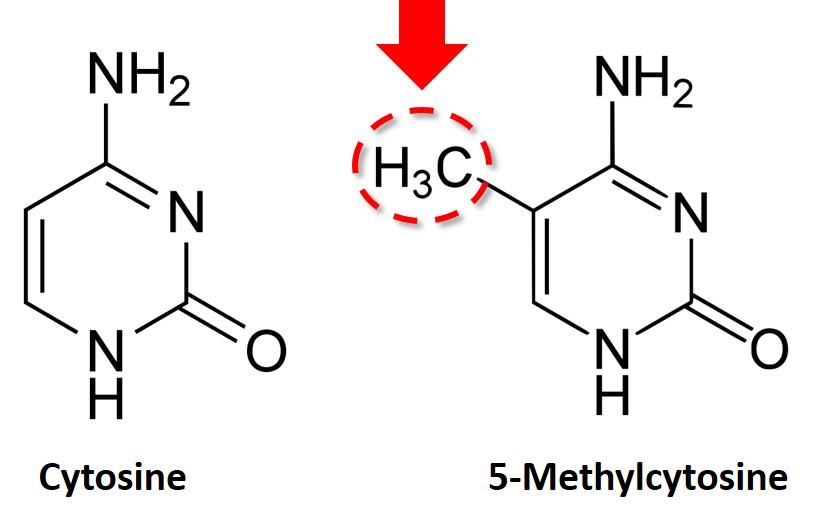 Bisulfite treatment of DNA
Bisulfite treatment of DNA
To be able to quantify DNA methylation levels at CpG sites using pyrosequencing, the DNA sample must be treated with sodium bisulfite. Bisulfite conversion is easily achieved by using commercially available kits.
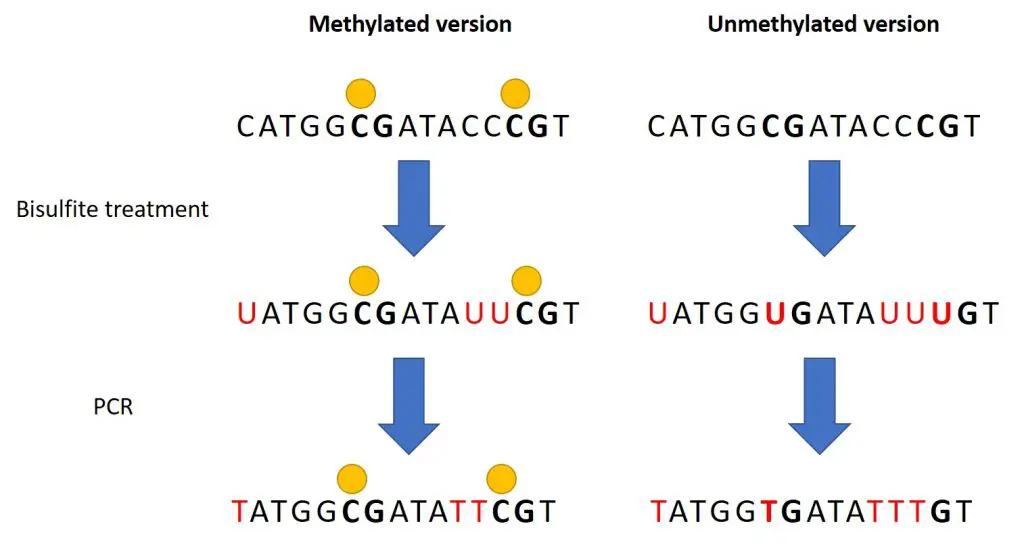 What the bisulfite treatment does is to change any unmethylated C’s into uracil (U) bases. This includes any C’s within and outside of CpG sites. Those C bases which are methylated (yellow circle in the above image) are resistant to this change and remain as a C after bisulfite treatment.
What the bisulfite treatment does is to change any unmethylated C’s into uracil (U) bases. This includes any C’s within and outside of CpG sites. Those C bases which are methylated (yellow circle in the above image) are resistant to this change and remain as a C after bisulfite treatment.
Next, following the PCR to amplify the DNA region of interest, any U base will change to the complementary thymine (T) base.
Calculating DNA methylation at CpG sites
Following the PCR reaction, samples can then be sequenced to determine the methylation content, usually presented as a percentage, of each CpG site. This can be achieved by numerous methods including pyrosequencing. The term bisulfite pyrosequencing refers to the sequencing of bisulfite-treated DNA.
The calculation of methylation at CpG sites is simply a ratio of C to T bases. For example, let’s say we bisulfite-treated our DNA and performed a PCR to amplify the region we want to sequence. Let’s pretend after PCR we got 5 amplicons (obviously you will get thousands after PCR – bear with me).
By looking at the image below you can see there are two CpG sites. The one to the left has three methylated amplicons and two unmethylated amplicons. Therefore, the methylation at this point will be 60% (3/5 x 100 = 60%). Now by looking at the CpG site on the right side, you can see that all of the amplicons are methylated therefore the methylation at this point will be 100% (5/5 x 100 = 100%). Simply, this is how bisulfite-pyrosequencing works to calculate DNA methylation percentages at individual CpG sites.
 Results from bisulfite pyrosequencing
Results from bisulfite pyrosequencing
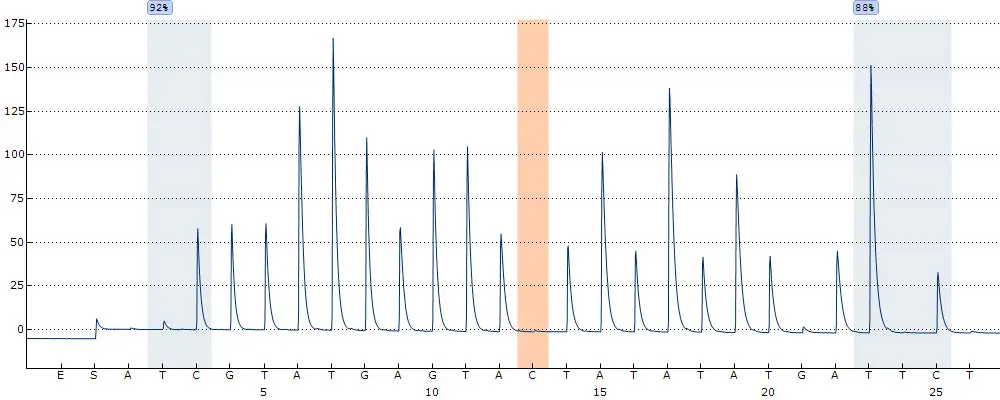
When looking at results from pyrosequencing, these are often presented as a pyrogram. A pyrogram reports the signal intensity following the addition of a base (y-axis) and the bases deposited (x-axis). The pyrogram (generated using the Qiagen PyroMark Q24 pyrosequencer) below contains two CpG sites highlighted in blue, with the corresponding DNA methylation percentage above each site.
 I have also included an annotated version of the pyrogram so you can see the different aspects of the graph.
I have also included an annotated version of the pyrogram so you can see the different aspects of the graph.
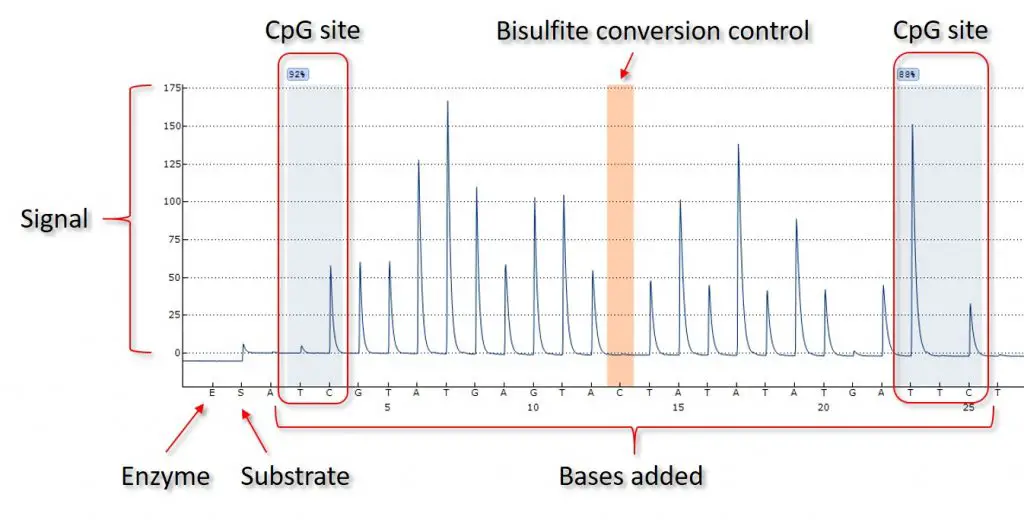 The PyroMark Q24 sequencer is also able to include bisulfite conversion quality control (highlighted in orange). The sequencer adds a C base to the reaction when C base is expected in the DNA sequence. Since the DNA has been bisulfite-treated, the C should not be incorporated (it should now be a T). However, if the bisulfite-treatment of the original DNA is not complete, then some signal will be gained here. This would indicate that some of the DNA was not fully converted. If so, the results within the CpG sites may not be accurate and the bisulfite treatment should be repeated.
The PyroMark Q24 sequencer is also able to include bisulfite conversion quality control (highlighted in orange). The sequencer adds a C base to the reaction when C base is expected in the DNA sequence. Since the DNA has been bisulfite-treated, the C should not be incorporated (it should now be a T). However, if the bisulfite-treatment of the original DNA is not complete, then some signal will be gained here. This would indicate that some of the DNA was not fully converted. If so, the results within the CpG sites may not be accurate and the bisulfite treatment should be repeated.
In sum, I have explained how bisulfite-pyrosequencing works to determine the DNA methylation content at CpG sites.




